|
|
| Since the early 1980s, geneticists have made considerable progress in using
genetic engineering techniques which are now applied in a variety of life
forms including microorganisms, plants, and animals. Genetic engineering
applications in breed improvement and pharmaceutical production are examples
of how the techniques improve our daily life. A considerable number of genes are responsible for the creation of life. Even a bacterium, one of the simplest life forms, possesses thousands of genes, and as for a human being, tens of thousands of genes work coordinately, although, the function of many of the genes is yet to be known. Given such facts, "improvement of a bacterium" for the sake of human benefit, cannot easily be achieved by manipulating just one or two genes. "Genome engineering technology" is next generation technology to theoretically improve genomes by manipulating tens/hundreds of genes on the basis of genome information. Aiming to develop the genome engineering technology targeting Corynebacterium, our group analyzed the genome structure of this bacterium by identifying the non-essential genes. We have so far successfully developed new genome engineering technologies such as the "large-scale genome deletion method" (that deletes tens of genes from the cell simultaneously), "multiple gene insertion method" (that inserts multiple genes simultaneously into the cell), and "makerless large-scale gene deletion/insertion method" (that enables gene manipulation without using antibiotics). We will continuously work on improvement of these technologies as well as development of new technologies. |
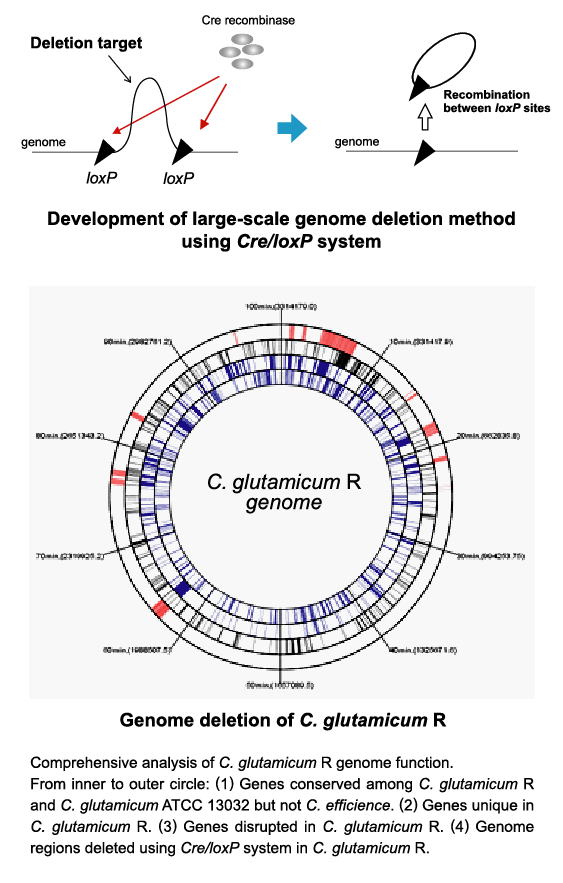
| Genome engineering technology for Corynebacterium (large-scale genome deletion) |
 |
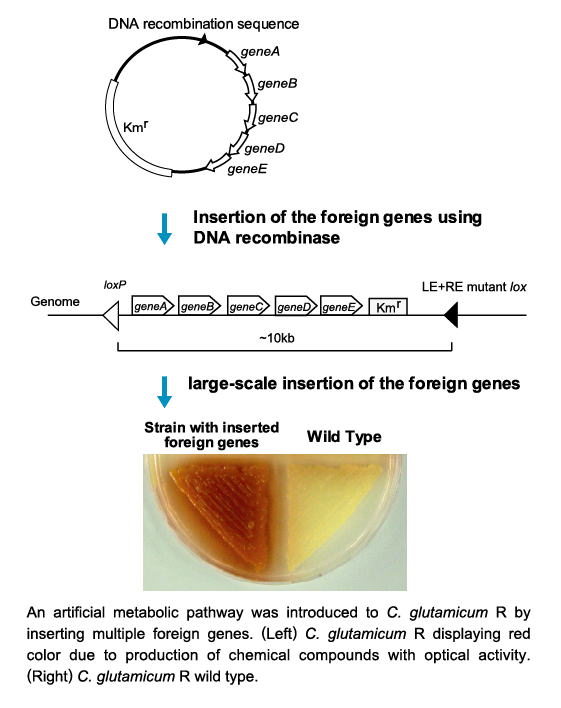
| Genome engineering technology for Corynebacterium (multiple-insertion of the foreign genes) |
 |
|
< References > Random segment deletion based on IS31831 and Cre/loxP excision system in Corynebacterium glutamicum. Appl. Microbiol. Biotechnol. 74: 1333-1341. 2007. Y. Tsuge, N. Suzuki, M. Inui and H. Yukawa. High-throughput transposon mutagenesis of Corynebacterium glutamicumand construction of a single-gene disruptant mutant library. Appl. Environ. Microbiol. 72: 3750-3755. 2006. N. Suzuki, N. Okai, H. Nonaka, Y. Tsuge, M. Inui and H. Yukawa. New multiple-deletion method for Corynebacterium glutamicum genome, using mutant lox sequence. Appl. Environ. Microbiol. 71: 8472-8480. 2005. N. Suzuki, H. Nonaka, Y. Tsuge, M. Inui and H. Yukawa. Manipulating corynebacteria, from individual genes to chromosomes. Appl. Environ. Microbiol. 71: 7633-7642. 2005. A.A. Vertès, M. Inui and H. Yukawa. Multiple large segment deletion method for Corynebacterium glutamicum. Appl. Microbiol. Biotechnol. 69: 151-161. 2005. N. Suzuki, H. Nonaka, Y. Tsuge, S. Okayama, M. Inui and H. Yukawa. Large-scale engineering of the Corynebacterium glutamicum genome. Appl. Environ. Microbiol. 71: 3369-3372. 2005. N. Suzuki, S. Okayama, H. Nonaka, Y. Tsuge, M. Inui and H. Yukawa. A new insertion sequence, IS14999, from Corynebacterium glutamicum. Microbiology 151: 501-508. 2005. Y. Tsuge, K. Ninomiya, N. Suzuki, M. Inui and H. Yukawa. Cre/loxP-mediated deletion system for large genome rearrangements in Corynebacterium glutamicum. Appl. Microbiol. Biotechnol. 67: 225-233. 2005. N. Suzuki, Y. Tsuge, M. Inui and H. Yukawa. Isolation and characterization of a native composite transposon, Tn14751, carrying 17.4 kilobases of Corynebacterium glutamicum chromosomal DNA. Appl. Environ. Microbiol. 71: 407-416. 2005. M. Inui, Y. Tsuge, N. Suzuki, A.A. Vertès and H. Yukawa. |